Long-read sequencing technologies, such as PacBio and Oxford Nanopore, have revolutionized genomics research by enabling the assembly of complex genomes, resolving structural variations, and improving haplotype phasing. However, the success of long-read sequencing heavily depends on the quality and integrity of DNA input. High molecular weight (HMW) DNA isolation is a crucial step that determines sequencing efficiency, read lengths, and overall data quality. In this article, we explore best practices for isolating high molecular weight DNA and optimizing it for long-read sequencing applications.
Why High Molecular Weight DNA Matters
Long-read sequencing platforms require intact, high-quality DNA to generate long sequencing reads, which provide superior genome assemblies compared to short-read technologies. Fragmented or degraded DNA results in shorter reads, reducing the ability to resolve structural variants and leading to incomplete assemblies.
Key benefits of HMW DNA for long-read sequencing:
● Enables reads of 10 kb to >100 kb for more comprehensive genome analysis
● Improves structural variant detection
● Enhances de novo genome assembly
● Provides better phasing of haplotypes
Challenges in Isolating High Molecular Weight DNA
Despite the benefits, isolating HMW DNA can be challenging. Factors that contribute to DNA degradation or shearing include:
● Harsh mechanical lysis or excessive pipetting
● Poor sample storage conditions
● Incomplete removal of proteins and contaminants
● Use of purification methods that fragment DNA
Addressing these challenges requires careful handling, optimized protocols, and the right reagents to maintain the integrity of high molecular weight DNA.
Best Practices for High Molecular Weight DNA Isolation
1. Choose the Right Sample Collection and Storage Methods
The integrity of DNA begins with proper sample handling. Fresh or properly preserved biological samples yield the best-quality DNA. Consider the following guidelines:
● Use fresh or flash-frozen samples to prevent degradation.
● Avoid excessive freeze-thaw cycles that can shear DNA.
● Store tissue samples in stabilizing solutions that protect nucleic acids.
● For blood samples, use anticoagulants that do not interfere with DNA integrity, such as EDTA or citrate.
2. Use a Gentle Lysis Method
Mechanical and enzymatic lysis methods should minimize DNA shearing. Common approaches include:
● Enzymatic digestion: Proteases and lysozymes gently break down cell walls without causing mechanical damage.
● Osmotic lysis: Used for certain cell types to avoid physical stress.
● Mechanical lysis (optional): If required, use a low-speed homogenization method to reduce shear forces.
● Nuclei isolation: Some protocols recommend isolating nuclei before DNA extraction to preserve ultra-long fragments.
3. Optimize DNA Purification for Long Fragment Recovery
Traditional column-based methods often shear DNA, making magnetic bead-based approaches preferable. Magnetic beads provide a gentle yet effective purification method that preserves long DNA fragments while removing contaminants like proteins and polysaccharides.
For best results:
● Use high-quality SPRI bead-based purification kits designed for HMW DNA recovery.
● Avoid vortexing or excessive pipetting, which can shear DNA.
● Ensure proper binding and elution conditions to retain long fragments.
● Select the appropriate binding buffer to maintain high DNA integrity.
4. Minimize Shearing During Handling
High molecular weight DNA is extremely susceptible to mechanical shearing. Follow these precautions:
● Use wide-bore pipette tips to reduce shear forces.
● Pipette slowly and gently to avoid breakage.
● Store DNA at 4°C to maintain integrity for short-term use.
● Use specialized storage solutions to prevent fragmentation over time.
● Avoid excessive mixing; use gentle inversion instead of vortexing.
5. Verify DNA Quality Before Sequencing
Assessing DNA quality is critical before proceeding with library preparation. Key metrics include:
● Size distribution: Use pulsed-field gel electrophoresis (PFGE) or capillary electrophoresis to confirm high molecular weight.
● Purity: Measure A260/A280 and A260/A230 ratios using a spectrophotometer.
● Concentration: Use Qubit or fluorometric methods to accurately quantify DNA.
● Integrity checks: Run agarose gel electrophoresis to visualize intact DNA.
Selecting the Best DNA Isolation Kits for Long-Read Sequencing
Choosing the right DNA isolation kit is essential for preserving high molecular weight DNA. MagBio Genomics offers industry-leading solutions optimized for long-read sequencing applications.
● HighPrep HMW DNA Kit: Designed for gentle and efficient isolation of HMW DNA from a variety of samples, minimizing shearing and maximizing yield.
● SPRI bead-based purification solutions: Ideal for cleanup and size selection of long DNA fragments.
● High Molecular Weight DNA Extraction Kits: Specifically formulated for PacBio Long Read Sequencing and Oxford Nanopore workflows.
Troubleshooting Common HMW DNA Isolation Issues
Even with optimized protocols, researchers may encounter challenges in HMW DNA isolation. Here are some common issues and their solutions:
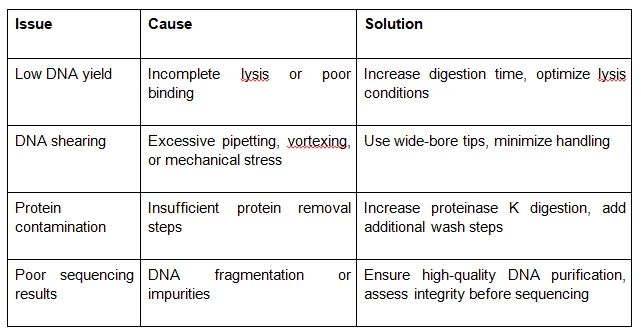
Long-Read Sequencing Workflow: Where HMW DNA Isolation Fits
The workflow for long-read sequencing typically includes:
- Sample collection and preservation
- HMW DNA isolation (critical step)
- DNA quality control and quantification
- Library preparation
- Sequencing on PacBio/Oxford Nanopore platforms
- Bioinformatics analysis
Any suboptimal step in this process can impact sequencing performance. Ensuring high-quality HMW DNA at the beginning improves the overall success rate of long-read sequencing experiments.
Maximize Your Long-Read Sequencing Success
Ensuring the highest quality HMW DNA is the foundation of successful long-read sequencing. By following best practices in sample handling, DNA isolation, purification, and quality assessment, researchers can achieve longer reads, better genome assemblies, and more accurate structural variant detection.
For researchers looking for High Molecular Weight DNA Isolation solutions, MagBio Genomics offers optimized magnetic bead-based kits designed to preserve high molecular weight DNA for long-read sequencing. HighPrep PCR is a paramagnetic bead-based post PCR clean-up reagent designed for efficient DNA purification and consistent DNA fragment size selection for NGS library preparation DNA cleanup. Learn more and explore our solutions at MagBio Genomics.













Sign in to leave a comment.