In the expanding field of genomics, high-quality DNA is the foundation of every successful experiment. For applications such as PacBio long-read sequencing, obtaining High Molecular Weight (HMW) DNA isn't just a bonus—it's a necessity. Long-read sequencing technologies demand ultra-pure, intact DNA fragments to deliver accurate, high-resolution genomic insights. This article delves deep into the techniques, challenges, and solutions related to isolating HMW DNA for PacBio sequencing, with a spotlight on DNA cleanup methods tailored for next-generation sequencing (NGS).
Why High Molecular Weight DNA Matters
When it comes to long-read sequencing, fragment length is everything. The PacBio HiFi sequencing platform excels in reading DNA fragments that are tens of kilobases in length. Unlike short-read technologies, which can manage fragmented DNA, long-read platforms require pristine DNA to produce accurate and comprehensive genomic data.
Benefits of HMW DNA in PacBio sequencing include:
● Longer read lengths, providing better genome assemblies.
● Reduced assembly gaps, particularly in repetitive regions.
● Improved detection of structural variants and haplotypes.
But getting there requires care. DNA can degrade or shear during isolation and purification, especially if you’re using methods not optimized for preserving long fragments.
Challenges in Isolating High Molecular Weight DNA
Despite advancements in extraction chemistry and tools, obtaining intact HMW DNA remains a challenge. Here are some common hurdles:
1. Mechanical Shearing
The most common cause of DNA fragmentation is mechanical force. Vortexing, pipetting, or even centrifugation can shear DNA. Once it's broken, there's no going back.
2. Enzymatic Degradation
Endogenous nucleases can chew up DNA during lysis if not properly inhibited. Maintaining a cold environment and using strong inhibitors is key.
3. Contaminants
Polysaccharides, proteins, and phenolic compounds can co-isolate with DNA, especially in plant and microbial samples. These inhibit downstream enzymatic reactions, like PCR and library construction.
4. Sample Type Variation
Different tissues yield DNA of varying quality and quantity. While blood and cultured cells are straightforward, plant tissues, fungi, and soil samples introduce complexity.
Best Practices for Isolating HMW DNA
Let’s walk through the process and highlight where you can maximize yield and integrity.
1. Sample Collection and Storage
Start strong. Immediately freeze samples in liquid nitrogen or dry ice if not processing right away. This preserves cellular structures and reduces enzymatic activity.
2. Gentle Cell Lysis
Use lysis buffers containing SDS, proteinase K, and RNase to gently break down membranes and proteins. Avoid vigorous mixing.
3. Phase Separation
Phenol-chloroform extraction is effective but harsh. Many labs now prefer magnetic bead-based methods, which are safer and scalable.
4. Precipitation and Resuspension
Isopropanol or ethanol can precipitate DNA. To minimize shearing, mix by gentle inversion. Let the DNA spool onto a glass rod or hook rather than centrifuging at high speeds.
5. Use of Magnetic Beads
Bead-based cleanups like those from MagBio Genomics use paramagnetic particles to bind DNA in a size-selective way. This helps retain large fragments while removing small debris and contaminants.
The Role of DNA Cleanup in Long-Read Sequencing
Even after extraction, your DNA needs to be cleaned and normalized. Contaminants can interfere with library preparation, reduce ligation efficiency, or even damage sequencing hardware.
What Cleanup Achieves:
● Removes inhibitors that affect polymerase activity.
● Eliminates small fragments that can dominate sequencing output.
● Improves ligation of sequencing adapters.
Choosing the Right Cleanup Kit for HMW DNA
The DNA cleanup step is just as critical as extraction. Kits like HighPrep™ PCR Clean-Up Kit from MagBio Genomics provide a robust solution:
Benefits of HighPrep™ PCR Clean-Up:
● Optimized for large DNA fragments.
● High recovery rates of DNA >10 kb.
● Scalable and automation-friendly.
● Cost-effective alternative to AMPure XP.
This kit utilizes Solid Phase Reversible Immobilization (SPRI) magnetic bead technology, allowing precise size selection and minimal sample loss.
PacBio Sequencing: Library Prep Considerations
Once you have clean, intact DNA, the next step is constructing your library.
Size Selection
Use pulsed-field gel electrophoresis or BluePippin to enrich for fragments above 15-20 kb. This ensures you’re making the most of PacBio’s capabilities.
DNA Damage Repair
Use a DNA Damage Repair mix to correct nicks and lesions. This enhances polymerase binding and sequencing accuracy.
Adapter Ligation
PacBio adapters require blunt-end ligation. Clean, intact DNA leads to higher ligation efficiency and better sequencing outcomes.
Troubleshooting HMW DNA Extraction
Even experienced researchers can hit roadblocks. Here are some common problems and solutions:
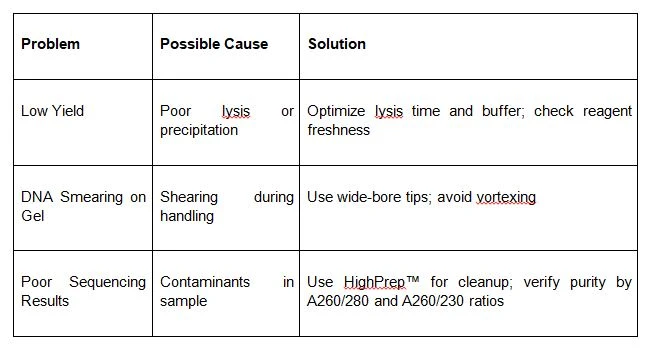
Real-World Applications
PacBio long-read sequencing is transforming genomics research:
● De novo genome assemblies: For organisms with no reference genome.
● Metagenomics: Resolving microbial communities and plasmids.
● Structural variant analysis: Detecting insertions, deletions, and translocations.
● Haplotype phasing: Long reads span multiple variants, making phasing easier.
But none of this is possible without clean, high-integrity DNA.
Future Trends in DNA Isolation
Automation and miniaturization are gaining momentum. Robotic systems integrated with magnetic bead-based kits allow high-throughput processing with minimal hands-on time. Additionally, researchers are developing non-invasive sampling and extraction methods to work with limited or degraded samples, such as ancient DNA.
Another trend? Multi-omics. Platforms are emerging to analyze DNA, RNA, and proteins from the same sample. This increases demand for universal, high-yield extraction protocols.
Conclusion
In the world of long-read sequencing, DNA quality is king. PacBio sequencing demands high molecular weight DNA that is free of contaminants and degradation. Achieving this level of quality requires careful attention to every step of the process, from sample collection to final cleanup.
Tools like MagBio Genomics' HighPrep™ PCR Clean-Up Kit make it easier than ever to purify DNA without sacrificing fragment size. Whether you're building a human genome from scratch or exploring microbial diversity in environmental samples, high-quality DNA is your starting point.
Get Started with HighPrep™ Today
Don't let subpar DNA compromise your sequencing. Discover how MagBio Genomics' HighPrep™ PCR Clean-Up Kit can help you purify and preserve your long DNA fragments for powerful, accurate PacBio sequencing.
Explore our High Molecular Weight DNA Isolation solutions today!

















Sign in to leave a comment.